Iterative proportional fitting of an abundance table to Hill-N2 marginals
Source:R/ipfN2marginals.R
ipf2N2.RdFunction for pre-processing/transforming an abundance table
by iterative proportional fitting,
so that the transformed table has marginals
proportional to N2 or N2(1-N2/N)
with N the number of elements in the margin.
Hill-N2 is the effective number of species. It is of intrinsic interest in
weighted averaging (CWM and SNC) as their variance is approximately
inversely proportional to N2 (ter Braak 2019),
and therefore of interest in dc_CA.
Arguments
- Y
abundance table (matrix or dataframe-like), ideally, with names for rows and columns.
- max_iter
maximum number of iterative proportional fitting (ipf) iterations. If
max_iter = 0, the columns are divided by their effective number or informativeness (N2orN2(1-N2/N), depending on the setting ofN2N_N2_species) without further pre-processing and the row sums are then, withN2N_N2_species = TRUE, sums of informativeness instead of effective number of informative species.- updateN2
logical, default
TRUE. IfFALSEthe marginal sums are proportional to the N2-marginals of the initial table, but the N2-marginals of the returned matrix may not be equal to their marginal sum. IfupdateN2 = TRUEandN2N_N2_species=TRUE(the default), the column marginals areN2(N-N2)/NwithNthe number of sites. The row sums are then proportional to, what we term, the effective number of informative species. IfN2N_N2_species = FALSE, the returned transformed table has N2 columns marginals, i.e.colSums(Y2) = N2species(Y2)withY2the return value ofipf2N2. If converged, N2 row marginals are equal to the row sums, i.e.rowSums(Y2) = approx. const*N2sites(Y2)andconsta constant.- N2N_N2_species
Set species marginal to the value of
N2(1-N2/N)for each species. DefaultTRUE. IfFALSE, the marginal is set to theN2value of each species.- N2N_N2_sites
Default
FALSEsets the marginal proportional to theN2value of each site. IfTRUE, the marginal is set toN2(1-N2/m), withmthe number of species.
Details
Applying ipf2N2 with N2N_N2_species=FALSE
to an presence-absence data table returns the same table.
However, a species that occurs everywhere (or in most of the sites)
is not very informative. This is acknowledged with the default option
N2N_N2_species=TRUE. Then,
with N2N_N2_species=TRUE, species that occur
in more than halve the number of sites are down-weighted, so that
the row sum is no longer equal to the richness of the site (the number of species),
but proportional to the number of informative species.
The returned matrix has the intended species marginal (column sums),
by construction of the algorithm, even without convergence.
On convergence, it has the intended site marginal (row sums).
References
ter Braak, C.J.F. (2019). New robust weighted averaging- and model-based methods for assessing trait-environment relationships. Methods in Ecology and Evolution, 10 (11), 1962-1971. doi:10.1111/2041-210X.13278
ter Braak, C.J.F. (2026). Fourth-corner latent variable models overstate confidence in trait–environment relationships and what to use instead Environmental and Ecological Statistics. doi:10.1007/s10651-025-00696-0
Examples
data("dune_trait_env")
# rownames are carried forward in results
rownames(dune_trait_env$comm) <- dune_trait_env$comm$Sites
Y <- dune_trait_env$comm[, -1] # must delete "Sites"
Y_N2 <- ipf2N2(Y, updateN2 = FALSE, N2N_N2_species = FALSE)
attr(Y_N2, "iter") # 61
#> [1] 61
# show that column margins of the transform matrix are
# equal to the Hill N2 values
diff(range(colSums(Y_N2) / apply(X = Y, MARGIN = 2, FUN = fN2))) # 4.440892e-16
#> [1] 4.440892e-16
diff(range(rowSums(Y_N2) / apply(X = Y, MARGIN = 1, FUN = fN2))) # 0.07077207
#> [1] 8.881784e-16
Y_N2i <- ipf2N2(Y, updateN2 = TRUE, N2N_N2_species = FALSE)
attr(Y_N2i, "iter") # 5
#> [1] 88
diff(range(colSums(Y_N2i) / apply(X = Y_N2i, MARGIN = 2, FUN = fN2))) # 2.220446e-15
#> [1] 7.771561e-16
diff(range(rowSums(Y_N2i) / apply(X = Y_N2i, MARGIN = 1, FUN = fN2))) # 8.881784e-16
#> [1] 1.332268e-15
# the default version:
Y_N2N_N2i <- ipf2N2(Y)
# ie.
# Y_N2N_N2i <- ipf2N2(Y, updateN2 = TRUE, N2N_N2_species = TRUE)
attr(Y_N2N_N2i, "iter") # 29
#> [1] 29
N2 <- apply(X = Y_N2N_N2i, MARGIN = 2, FUN = fN2)
N <- nrow(Y)
diff(range(colSums(Y_N2N_N2i) / (N2 * (N - N2)))) # 4.857226e-17
#> [1] 4.857226e-17
N2_sites <- apply(X = Y_N2N_N2i, MARGIN = 1, FUN = fN2)
R <- rowSums(Y_N2N_N2i)
N * max(N2_sites / sum(N2_sites) - R / sum(R)) # 0.006116092
#> [1] 0.006116092
sum(Y > 0)
#> [1] 179
sum(Y_N2N_N2i)
#> [1] 94.12255
sum(Y)
#> [1] 626
mod0 <- dc_CA(formulaEnv = ~ A1 + Moist + Mag + Use + Manure,
formulaTraits = ~ SLA + Height + LDMC + Seedmass + Lifespan,
response = Y,
dataEnv = dune_trait_env$envir,
dataTraits = dune_trait_env$traits,
divide = FALSE,
verbose = FALSE)
mod1 <- dc_CA(formulaEnv = ~ A1 + Moist + Mag + Use + Manure,
formulaTraits = ~ SLA + Height + LDMC + Seedmass + Lifespan,
response = Y_N2N_N2i,
dataEnv = dune_trait_env$envir,
dataTraits = dune_trait_env$traits,
verbose = FALSE)
#> Argument divideBySiteTotals set to FALSE, as species totals are proportional to N2(N-N2).
#> You can overrule this by specifying divideBySiteTotals explicitly.
mod1$eigenvalues / mod0$eigenvalues
#> dcCA1 dcCA2 dcCA3 dcCA4 dcCA5
#> 1.1829169 1.3376443 1.3744124 1.5197560 0.8780165
# ratios of eigenvalues greater than 1,
# indicate axes with higher (squared) fourth-corner correlation
# ipf2N2 for a presence-absence data matrix
Y_PA <- 1 * (Y > 0)
Y_PA_N2 <- ipf2N2(Y_PA, N2N_N2_species = FALSE)
attr(Y_PA_N2, "iter") # 3
#> [1] 3
diff(range(Y_PA - Y_PA_N2)) # 7.771561e-16, i.e no change
#> [1] 7.771561e-16
Y_PA_N2i <- ipf2N2(Y_PA, N2N_N2_species = TRUE)
attr(Y_PA_N2i, "iter") # 567
#> [1] 567
N_occ <- colSums(Y_PA) # number of occurrences of species
N <- nrow(Y_PA)
plot(N_occ, colSums(Y_PA_N2i))
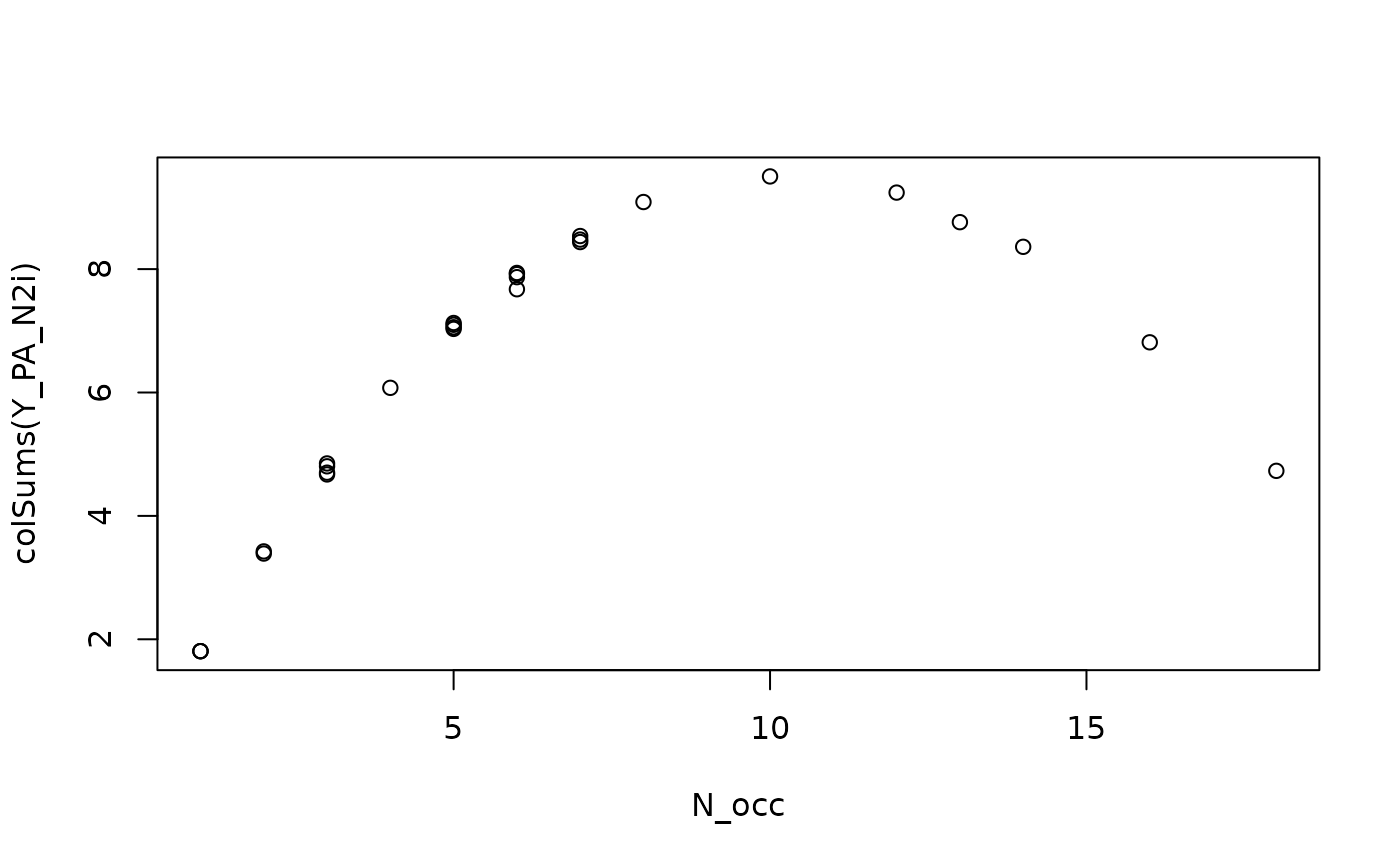 cor(colSums(Y_PA_N2i), N_occ * (N - N_occ)) # 0.9826123
#> [1] 0.9826123
mod2 <- dc_CA(formulaEnv = ~ A1 + Moist + Mag + Use + Manure,
formulaTraits = ~ SLA + Height + LDMC + Seedmass + Lifespan,
response = Y_PA,
dataEnv = dune_trait_env$envir,
dataTraits = dune_trait_env$traits,
divideBySiteTotals = FALSE,
verbose = FALSE)
mod3 <- dc_CA(formulaEnv = ~ A1 + Moist + Mag + Use + Manure,
formulaTraits = ~ SLA + Height + LDMC + Seedmass + Lifespan,
response = Y_PA_N2i,
dataEnv = dune_trait_env$envir,
dataTraits = dune_trait_env$traits,
verbose = FALSE)
#> Argument divideBySiteTotals set to FALSE, as species totals are proportional to N2(N-N2).
#> You can overrule this by specifying divideBySiteTotals explicitly.
mod3$eigenvalues / mod2$eigenvalues
#> dcCA1 dcCA2 dcCA3 dcCA4 dcCA5
#> 1.578066 1.309621 1.649829 1.648562 1.370179
# ratios of eigenvalues greater than 1,
# indicate axes with higher (squared) fourth-corner correlation
cor(colSums(Y_PA_N2i), N_occ * (N - N_occ)) # 0.9826123
#> [1] 0.9826123
mod2 <- dc_CA(formulaEnv = ~ A1 + Moist + Mag + Use + Manure,
formulaTraits = ~ SLA + Height + LDMC + Seedmass + Lifespan,
response = Y_PA,
dataEnv = dune_trait_env$envir,
dataTraits = dune_trait_env$traits,
divideBySiteTotals = FALSE,
verbose = FALSE)
mod3 <- dc_CA(formulaEnv = ~ A1 + Moist + Mag + Use + Manure,
formulaTraits = ~ SLA + Height + LDMC + Seedmass + Lifespan,
response = Y_PA_N2i,
dataEnv = dune_trait_env$envir,
dataTraits = dune_trait_env$traits,
verbose = FALSE)
#> Argument divideBySiteTotals set to FALSE, as species totals are proportional to N2(N-N2).
#> You can overrule this by specifying divideBySiteTotals explicitly.
mod3$eigenvalues / mod2$eigenvalues
#> dcCA1 dcCA2 dcCA3 dcCA4 dcCA5
#> 1.578066 1.309621 1.649829 1.648562 1.370179
# ratios of eigenvalues greater than 1,
# indicate axes with higher (squared) fourth-corner correlation